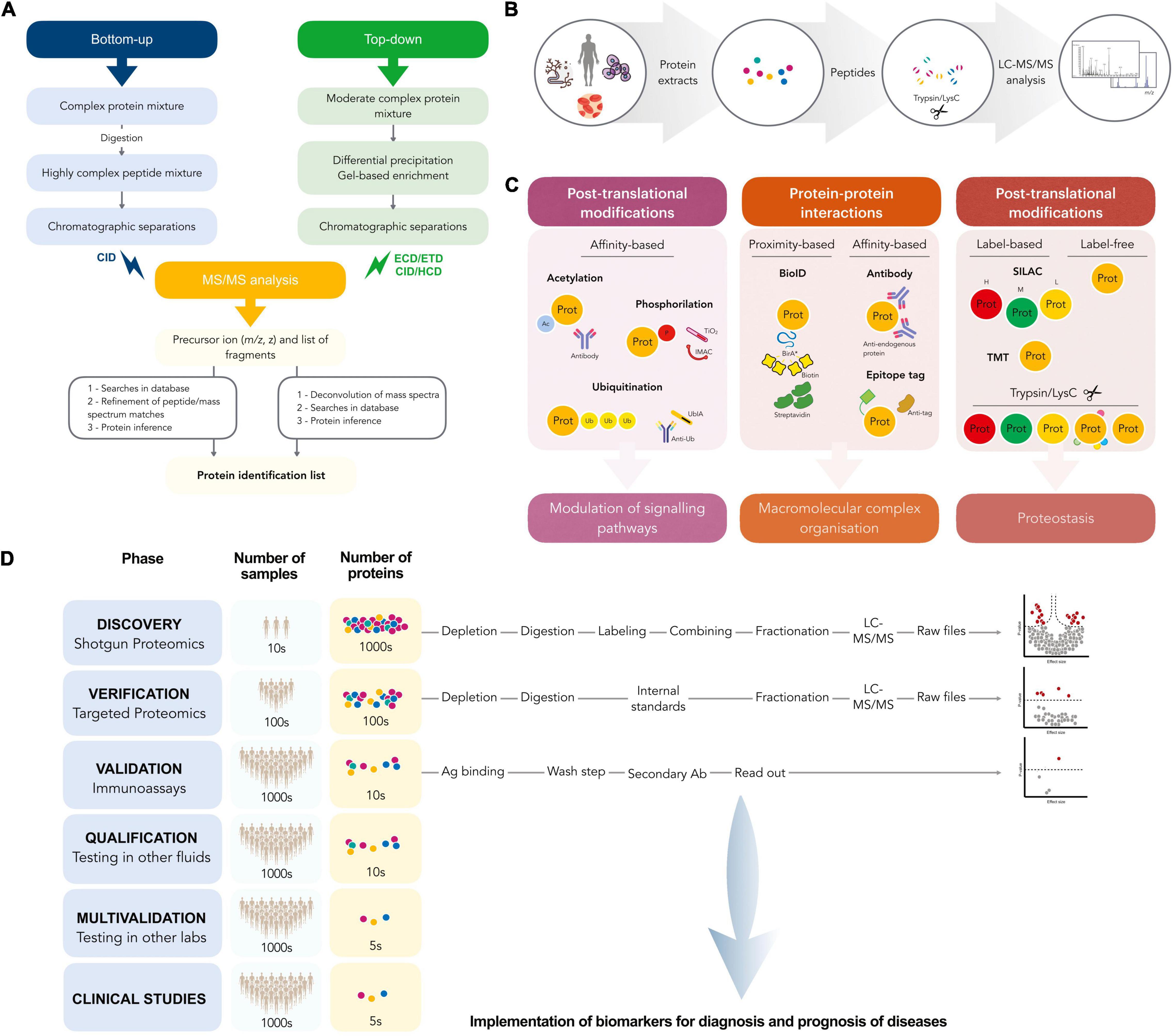
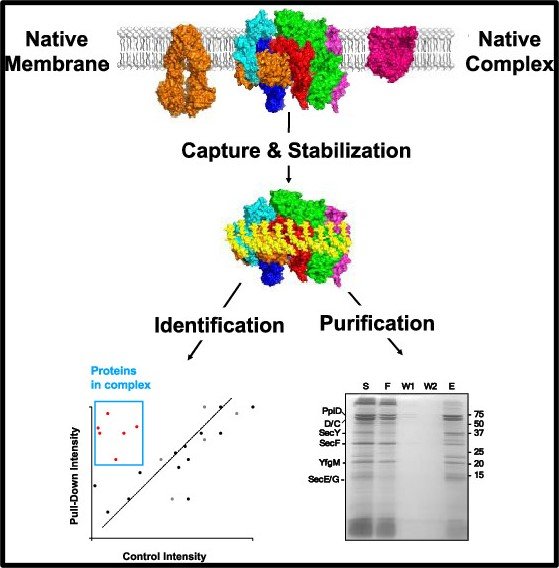
Quantitative Proteomics-Based Substrate Screening Revealed Cyclophilin Stabilization Regulated by Deubiquitinase Ubp7 | Journal of Proteome Research

prolfqua: A Comprehensive R-Package for Proteomics Differential Expression Analysis | Journal of Proteome Research

Proteomic Analysis of Integrin-Associated Complexes Identifies RCC2 as a Dual Regulator of Rac1 and Arf6 | Science Signaling

A proteomic approach reveals integrin activation state-dependent control of microtubule cortical targeting | Nature Communications

Proteomics and Beyond: Cell Decision-Making Shaped by Reactive Electrophiles: Trends in Biochemical Sciences

RIPA Lysis Buffer For Extraction And Solubilization Of Proteins From Cultured Mammalian Adherent And Suspension Cells, 125mL
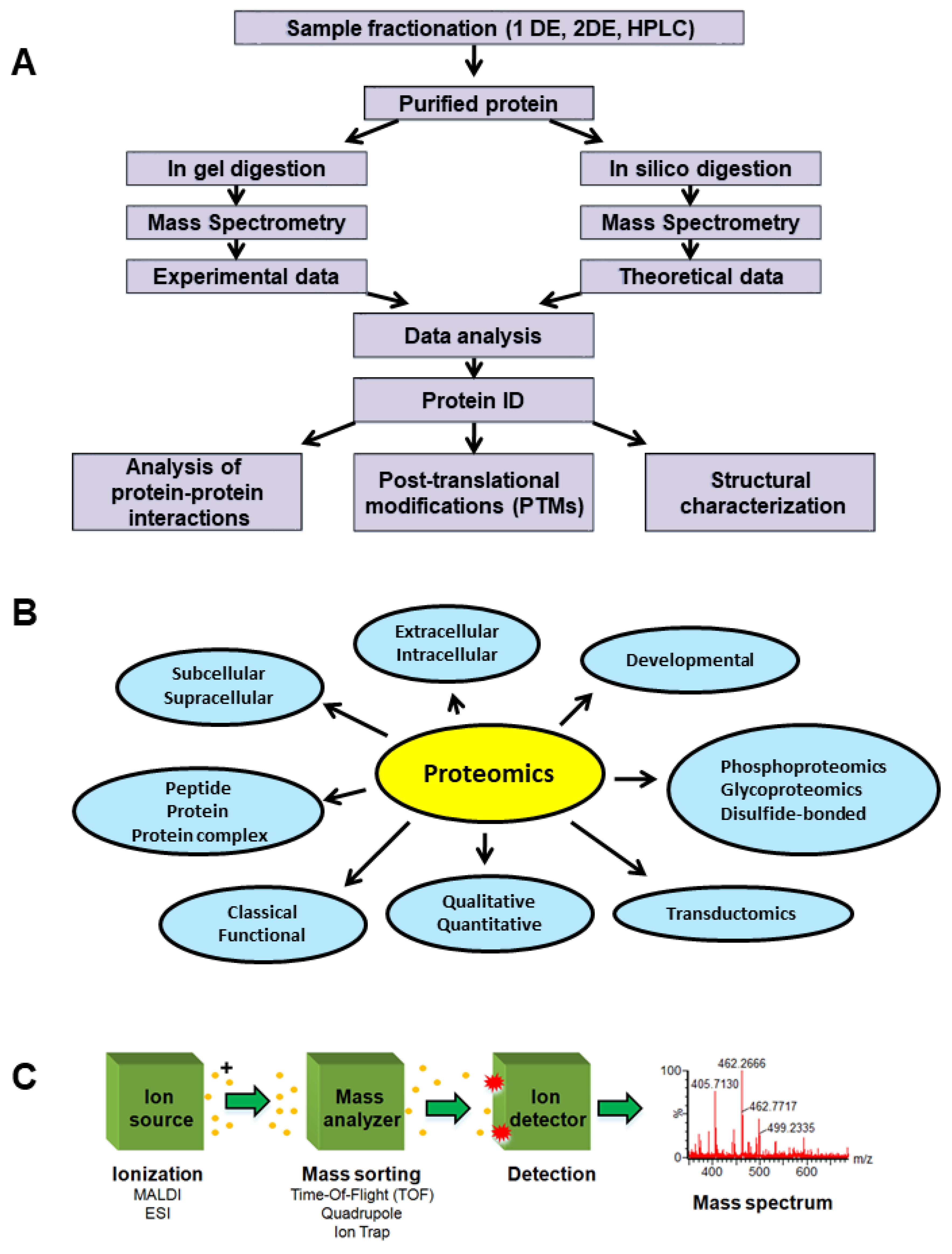
Proteomes | Free Full-Text | Proteomics-Based Identification of Dysregulated Proteins in Breast Cancer

Comparative Evaluation of MaxQuant and Proteome Discoverer MS1-Based Protein Quantification Tools | Journal of Proteome Research

Global Analysis of the Mitochondrial N-Proteome Identifies a Processing Peptidase Critical for Protein Stability: Cell
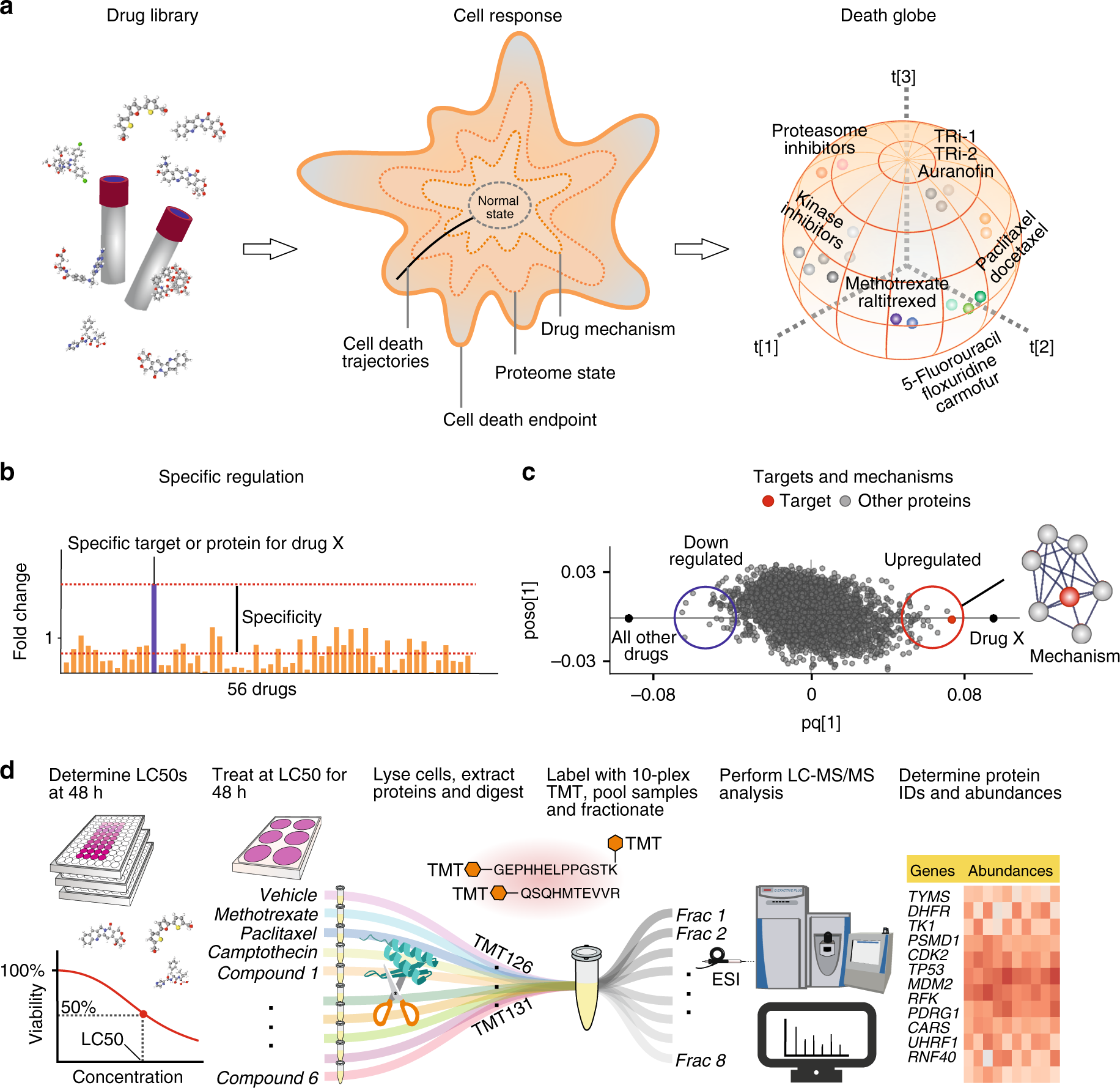
ProTargetMiner as a proteome signature library of anticancer molecules for functional discovery | Nature Communications

Contribution of Mass Spectrometry-Based Proteomics to the Understanding of TNF-α Signaling | Journal of Proteome Research
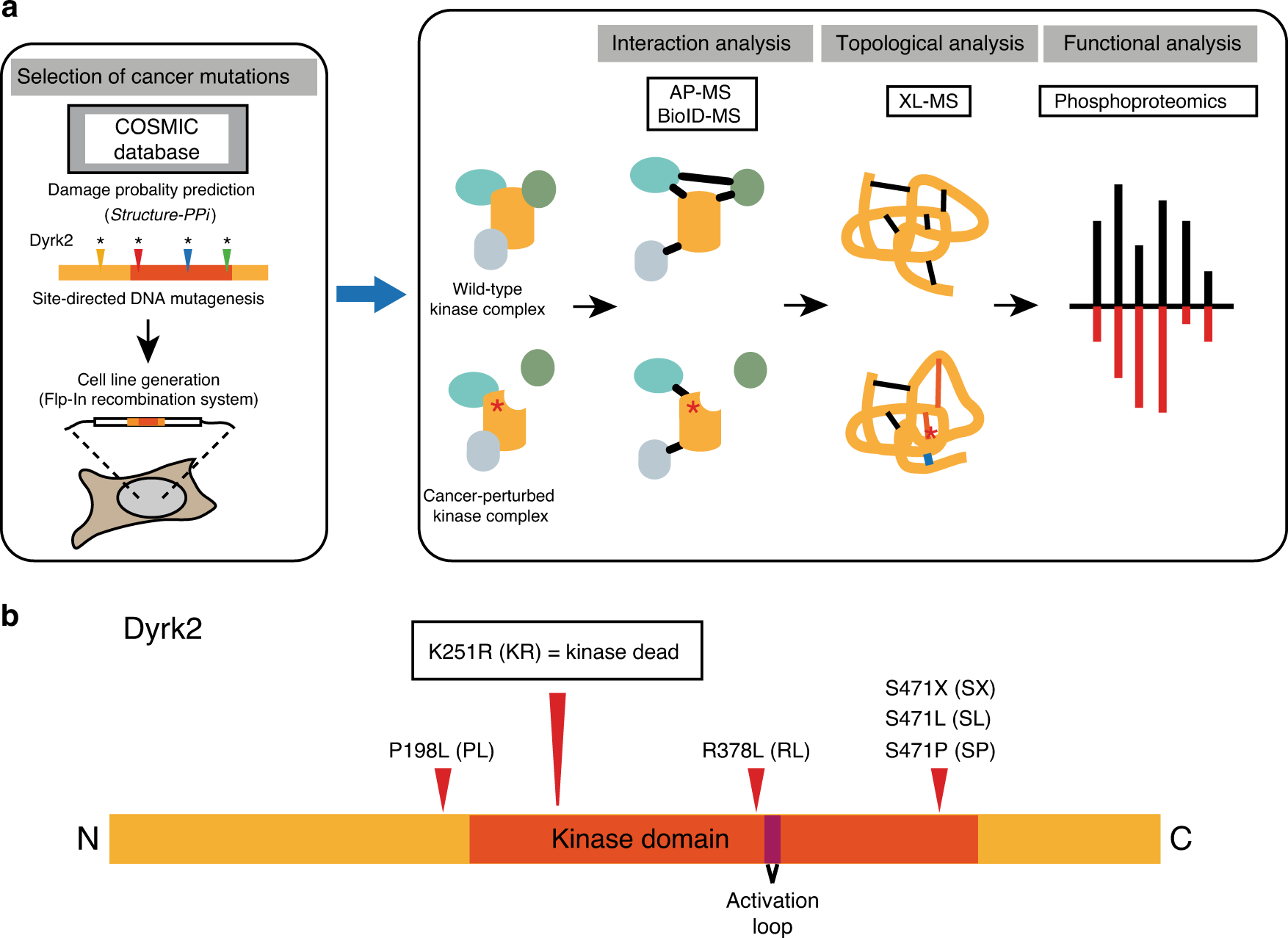
Multi-layered proteomic analyses decode compositional and functional effects of cancer mutations on kinase complexes | Nature Communications